Chapter 14: What We Don't Know Yet
Chapter 14: What We Don’t Know Yet
Part V: The Future
It was late on a Thursday when I closed my laptop for what I told myself would be the last time on this project. The final molecular dynamics trajectory had finished rendering. The last network visualization had been exported. Four studies, fifteen compounds, dozens of targets, hundreds of docking calculations, thousands of hours of computation — all distilled into the chapters you’ve been reading.
There was a cup of coffee beside me. There is always a cup of coffee beside me. It had gone cold, which is what happens when you spend forty minutes staring at binding energy histograms instead of drinking. I picked it up, looked at the dark surface, and a thought arrived that was equal parts humbling and electrifying:
After four papers and this entire book, how much do I actually know about what’s in this cup?
The honest answer is: more than I did before, and far less than I need to.
This chapter is the scientific conscience of everything that came before it. In thirteen chapters, I’ve walked you through computational predictions, network maps, molecular docking scores, and pharmacokinetic models. I’ve tried to be careful with language throughout — “predicted,” “suggested,” “modeled” — but I know how the human mind works. After enough pages of confident-sounding science, the caveats blur and what remains feels like certainty.
So let me be explicit, one last time, about the boundary between what our models generated and what biology has actually confirmed.
What Our Computational Models Can Tell Us
Let me start with what we did accomplish, because I don’t want intellectual honesty to shade into false modesty. The four studies that form the backbone of this book produced genuine, novel, testable contributions to coffee science.
We showed that coffee diterpenes — cafestol and kahweol — are predicted to bind the farnesoid X receptor with pharmaceutical-grade affinity. The docking scores we obtained for these compounds at FXR are in the same range as synthetic drugs designed specifically for that target. This is not a trivial finding. FXR is a master regulator of bile acid metabolism, lipid homeostasis, and glucose regulation. If those predicted affinities hold up experimentally, it would provide a molecular explanation for coffee’s well-documented but poorly understood effects on liver health and cholesterol metabolism.
We mapped coffee’s bioactive network — six key compounds interacting with ten validated targets across four major signaling pathways — and showed that it has organized multi-target activity. This is not a random scatter of weak interactions. The network has structure. Multiple compounds from different chemical families hit complementary targets in interconnected biological pathways. Computationally speaking, coffee looks less like a food and more like a carefully designed combination drug — except no one designed it.
We identified what I called the 75-125x Maillard bottleneck — the concentration factor required for roasting-generated melanoidins to reach predicted bioactive thresholds. This puts a number, for the first time, on why coffee’s development phase (the chemistry of roasting) matters so much for its biological activity.
We predicted that ten of our fifteen studied compounds can cross the blood-brain barrier. These predictions are based on each molecule’s size, how easily it dissolves in fat versus water (lipophilicity), how polar its surface is, and how many hydrogen bonds it can form. All fifteen compounds also pass Lipinski’s Rule of Five — a standard checklist indicating that a molecule has the physical properties needed to work as a drug.
These are real findings. They advance understanding. They generate specific, testable hypotheses.
But here is the sentence I need you to carry with you: not one of these predictions has been clinically validated. Every result in this book lives in the space between computation and confirmation.
What We Still Don’t Know: The Honest List
We Haven’t Confirmed the Docking Predictions Experimentally
Our molecular docking studies predicted that cafestol binds FXR with high affinity, and that specific binding poses orient the molecule in the receptor’s ligand-binding domain in ways that would activate transcription. These predictions were generated using validated software, established protocols, and well-characterized crystal structures.
But nobody — not my lab, not anyone else — has confirmed these specific interactions with X-ray crystallography of the cafestol-FXR complex, or with competitive binding assays, or with any direct biophysical measurement. The docking scores are calculated estimates. They are sophisticated estimates, generated by algorithms that have been validated against known crystal structures with good accuracy. But they are not measurements.
The difference between a predicted binding affinity and a measured one is the difference between a weather forecast and the actual weather. The forecast might be excellent. You should still look out the window. And right now, for cafestol at FXR, nobody has looked out the window.
We Don’t Know In Vivo Concentrations Where They Matter Most
Our blood-brain barrier analysis predicted that two-thirds of coffee’s bioactive compounds have the molecular properties needed to cross from blood into brain tissue. The physicochemical criteria we used — molecular weight below 450 daltons, adequate lipophilicity, limited hydrogen bonding — are well-established predictors.
But prediction of permeability is not measurement of concentration. To my knowledge, nobody has ever measured cafestol levels in human cerebrospinal fluid after coffee consumption. Nobody has quantified chlorogenic acid concentrations in specific brain regions. The pharmacokinetic models tell us these molecules could get there. Whether they do get there, and at what concentrations, and for how long — that remains unknown.
This matters enormously because biological activity is concentration-dependent. A compound that reaches the brain at nanomolar concentrations will behave very differently from one that arrives at micromolar levels. Our models don’t distinguish between these scenarios.
Melanoidin Structure Remains Incompletely Characterized
Melanoidins — the large, brown, heterogeneous polymers formed during coffee roasting through the Maillard reaction — are arguably the most important and least understood class of compounds in the cup. They represent up to 25% of dry coffee weight. They reach the colon largely intact and interact extensively with the gut microbiome.
And we still don’t fully know what they look like.
I’ve stared at melanoidin mass spectra that look nothing like the clean, sharp peaks you get for caffeine or chlorogenic acid. Instead: a broad, featureless hump — the analytical equivalent of asking “who’s in this room?” and getting the answer “people.” These polymers range from roughly 3 to over 100 kilodaltons in mass. No two molecules are identical. They incorporate amino acids, sugars, chlorogenic acids, and lipids into complex, branching architectures that resist the standard tools of structural chemistry.
In our studies, I modeled melanoidin interactions using representative fragments and substructures, not complete molecules. This was a necessary simplification — you cannot dock a molecule whose structure you don’t fully know. But it means our melanoidin predictions are, at best, approximations of approximations.
I confess that this particular unknown fascinates me more than it frustrates me. A quarter of what’s in your cup — the brown color, the body, the mouthfeel, the prebiotic activity — comes from molecules we cannot fully describe. There is something both humbling and wonderful about drinking a mystery every morning.
We Didn’t Model Synergistic Effects Between Compounds
Our network analysis mapped each compound to its predicted targets individually. Cafestol interacts with FXR. Chlorogenic acid interacts with NF-kB. Trigonelline interacts with its receptor set. The network shows how these individual interactions converge on shared pathways.
What it does not show is how these compounds interact with each other. In an actual cup of coffee, all fifteen bioactives coexist simultaneously. They may compete for the same binding sites. They may alter each other’s absorption, metabolism, or excretion. One compound might enhance another’s activity (synergy) or suppress it (antagonism).
Modeling these compound-compound interactions is computationally orders of magnitude harder than modeling compound-target interactions. Think of the difference between predicting how one guest at a dinner party will behave versus predicting the dynamics of all fifteen guests simultaneously — who talks over whom, who forms alliances, who cancels someone else out. We didn’t attempt it. Nobody has, for coffee, at the scale we would need.
If you’re keeping count, that’s three major unknowns — and I haven’t reached the one that keeps me up at night.
Dose-Response Remains an Open Question
Here is the gap that probably matters most to you, personally: we cannot tell you, from computation alone, how much coffee you need to drink for any of these predicted effects to manifest. Three cups? Five? One carefully prepared pour-over? We don’t know.
The journey from “this compound docks into this receptor in a computer simulation” to “drinking two cups of this particular coffee, prepared this particular way, produces this measurable health effect in this type of person” involves so many steps — absorption, metabolism, distribution, tissue-specific concentration, receptor occupancy kinetics, downstream signaling cascades, compensatory mechanisms — that our models address only the very first link in a very long chain.
Individual Variation Is Enormous and Unmodeled
Your genetics determine how fast you metabolize caffeine (via CYP1A2 polymorphisms), how you process chlorogenic acids, and how your bile acid receptors respond to diterpene binding. Your gut microbiome — which is as individual as a fingerprint — determines how melanoidins are fermented in your colon and what metabolites are produced. Your health status, your medications, your diet, your age — all of these modulate how coffee compounds behave in your specific body.
Our computational models describe average molecular behavior in idealized conditions. Your body is not an average, and its conditions are not ideal. I think about this every time someone asks me, “So is coffee good for me?” The honest answer — “it depends on your CYP1A2 genotype, your microbiome composition, your NF-kB baseline, and about forty other variables I can’t measure from here” — is not what anyone wants to hear over breakfast. But it’s the truth. The gap between population-level computational prediction and individual biological response is one of the largest unsolved problems not just in coffee science, but in all of pharmacology.
What you can do right now: You cannot sequence your microbiome over breakfast. But you can pay attention to your own body’s responses — and that personal data is more valuable than any computational model. Track your energy, your sleep, your digestion across different brewing methods and roast levels for a month. Note what time you drink, what you eat alongside it, how you feel two hours later. You are running an n=1 experiment whether you intend to or not. You might as well collect the data.
Every day, approximately 2.25 billion cups of coffee are consumed worldwide. That makes coffee one of the most consumed bioactive preparations on Earth — a complex mixture of hundreds of compounds with documented effects on the liver, brain, cardiovascular system, and gut microbiome, consumed daily by roughly a third of the planet’s adult population.
And yet, the scientific rigor applied to understanding coffee’s biological mechanisms is a fraction of what we apply to pharmaceutical compounds consumed by far fewer people. A single drug candidate will undergo years of binding assays, pharmacokinetic studies, animal models, and phased clinical trials before reaching patients. Coffee — consumed by billions, containing dozens of bioactives, interacting with scores of biological targets — has had almost none of this systematic characterization at the molecular level.
There is a profound scale mismatch between how much coffee we consume and how well we understand what it does inside us. This book is, in part, an argument that computational tools can begin to close that gap — not by replacing experimental science, but by telling experimentalists where to look first.
The NR4A1 Discovery: When the Network Surprises You
I want to spend a moment on what I consider the single most interesting — and most uncertain — finding from our work.
When we constructed the compound-target interaction network for coffee bioactives, running our fifteen compounds against databases of known and predicted protein targets, most of what came back was expected. We saw the usual suspects: NF-kB for inflammation, PPARs for lipid metabolism, AChE for neurological effects. These are targets that the coffee literature has circled around for years.
But the network also returned NR4A1. And I sat up straighter in my chair.
NR4A1, also known as Nur77, is a nuclear receptor — a protein that sits in the cell nucleus and directly regulates gene expression. It’s involved in metabolic regulation, inflammatory responses, and cell survival decisions. It has been the subject of intense pharmaceutical interest for conditions ranging from metabolic syndrome to certain cancers.
It had not, to my knowledge, been previously connected to coffee bioactives.
This is the kind of finding that makes computational science both thrilling and treacherous. Thrilling, because the network identified a biologically plausible, pharmacologically significant target that nobody had thought to look at in the context of coffee. Treacherous, because the prediction is only as good as the databases and algorithms that generated it, and a novel prediction has, by definition, zero experimental validation.
I want to be precise about what I can and cannot claim: I cannot tell you that coffee affects NR4A1 in your body. I can tell you that our computational models predict an interaction between specific coffee bioactives and this receptor that, based on the network topology, appears to be non-random and worth investigating. The distance between those two statements is exactly the distance between hypothesis and knowledge.
NR4A1, or Nur77, belongs to a family of “orphan” nuclear receptors — so named because, for years, scientists didn’t know what natural molecule activated them. Unlike estrogen receptors (activated by estrogen) or vitamin D receptors (activated by vitamin D), Nur77 seemed to operate without a clear physiological ligand.
What we do know: Nur77 acts as a molecular switch in metabolism and inflammation. When activated, it can shift cells from inflammatory to anti-inflammatory programs, influence glucose uptake, and modulate lipid processing. It sits at a crossroads of pathways that are individually well-studied in coffee research — but nobody had identified this particular crossroads as relevant to coffee.
Our network analysis flagged it because multiple coffee bioactives showed predicted interactions with NR4A1 or with proteins directly upstream of it in signaling cascades. The convergence of several independent compounds on the same previously unrecognized target is what made the prediction statistically notable.
Whether it is biologically real remains entirely open. But if validated, it would provide a unifying mechanism connecting coffee’s anti-inflammatory and metabolic effects through a single nuclear receptor — an elegant explanation for observations that currently lack a molecular home.
What Needs to Happen Next
I am a computational scientist. I generate hypotheses. The confirmation belongs to other kinds of laboratories — wet labs with their pipettes and centrifuges, their cell cultures smelling faintly of warm plastic, their binding assays and mass spectrometers and, eventually, clinical research facilities with human participants. But I can describe what I believe the research agenda should look like based on what our studies revealed.
First, experimental binding assays. The docking predictions for cafestol and kahweol at FXR are specific enough to be directly testable. Isothermal titration calorimetry, surface plasmon resonance, or fluorescence-based competitive binding assays could confirm or refute our predicted affinities within months. These are not exotic techniques. They are routine in any well-equipped biochemistry department. The fact that they haven’t been applied to coffee diterpenes at FXR reflects priorities, not technical barriers.
Second, pharmacokinetic measurements. We need actual concentration-time profiles of coffee bioactives in human blood and tissues after coffee consumption. Not just caffeine — we have those data. Cafestol, kahweol, trigonelline, N-methylpyridinium, chlorogenic acid metabolites. In plasma, certainly. In cerebrospinal fluid, ideally. Until we know what concentrations these compounds actually reach in target tissues, our permeability and distribution predictions float in a vacuum.
Third, better melanoidin characterization. This is perhaps the hardest technical challenge on the list. Melanoidins resist standard structural analysis because of their heterogeneity and size. But analytical chemistry is advancing — high-resolution mass spectrometry, ion mobility spectrometry, cryo-electron microscopy for large complexes. I believe that within the coming years, we will have substantially better structural models for at least the dominant melanoidin species in coffee. When we do, the computational analysis will need to be revisited with realistic structures rather than fragments.
Fourth, study designs that match coffee’s complexity. Most clinical studies of coffee treat it as a single exposure — “coffee” versus “no coffee” — and measure single outcomes. Our network analysis shows that coffee operates through multi-compound, multi-target, multi-pathway mechanisms. Future clinical studies should be designed to capture this complexity: measuring multiple biomarkers simultaneously, stratifying by genetic polymorphisms in relevant metabolic enzymes, and accounting for preparation method (which determines diterpene content, as we discussed in earlier chapters).
While we wait for science to close these gaps, here is what you can act on today, based on what the evidence already supports:
If cholesterol concerns you: Use a paper filter. This is the single most evidence-backed brewing decision in all of coffee science (Chapter 2).
If you want maximum polyphenols: Choose lighter roasts. CGA content drops from 80% to as low as 5% from light to dark roast (Chapter 1).
If coffee disrupts your sleep: Your CYP1A2 genotype likely makes you a slow metabolizer. Set a personal caffeine cutoff time and stick to it (Chapter 6).
If gut comfort matters: Darker roasts may be gentler, thanks to higher NMP content and lower CGA acidity (Chapter 1).
If you want the most from coffee’s predicted network effects: Consistency matters more than quantity. The epidemiological associations are strongest at 3-5 cups per day, consumed regularly (Chapter 7).
The Vision: What Computational Coffee Science Could Become
I started this project as a physicist who had wandered into pharmaceutical science and then looked sideways at the cup on her desk. The tools I used — molecular docking, network pharmacology, ADMET prediction, systems biology — were developed for drug discovery. They were designed to evaluate synthetic compounds aimed at single targets for specific diseases.
Applying them to coffee was, in some ways, an act of creative misuse. Coffee is not a drug. It is not designed to treat anything. It is a complex natural extract consumed for pleasure, for culture, for the ritual of preparation, for the warmth of the cup in your hands on a cold morning. Nobody needs a docking score to enjoy their espresso.
But 2.25 billion cups a day represents one of the largest uncontrolled biological experiments in human history. Every morning, before the sun has fully risen in some time zone, millions of hands are reaching for kettles, pressing buttons on espresso machines, pouring water over grounds in ceramic drippers. Each one of those rituals delivers a complex mixture of bioactive compounds into a unique human body — compounds that interact with nuclear receptors, inflammatory pathways, neurotransmitter systems, and gut microbiota. We owe it to those billions of daily consumers — we owe it to ourselves — to understand what those interactions are, with a rigor that matches the scale.
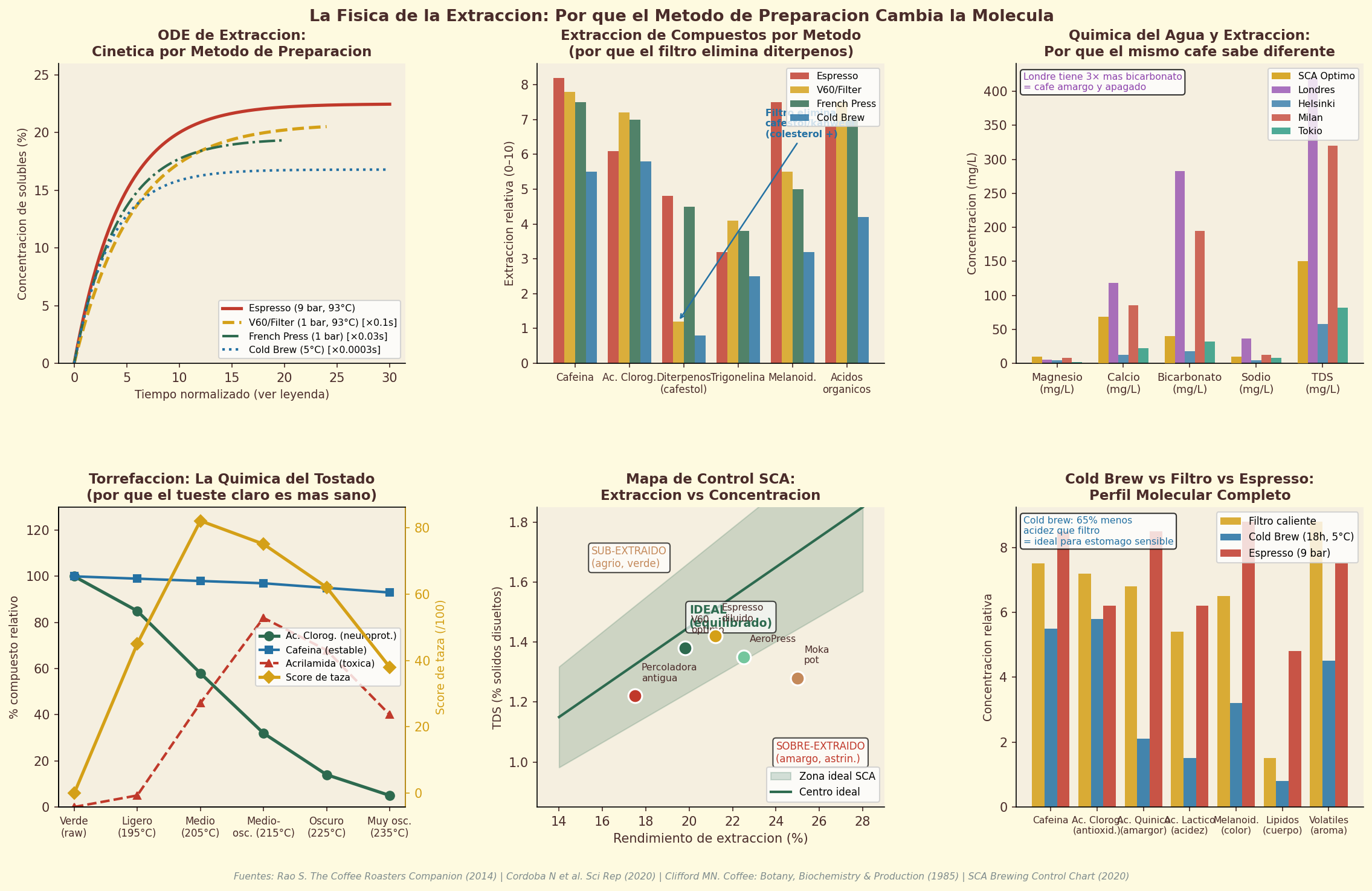
Figure 14.1. The coffee extraction process: how different brewing methods extract different molecular profiles from the same roasted beans, shaping the bioactive composition of every cup.
That is what I hope computational coffee science can become: a field that bridges food science and drug discovery, using the powerful predictive tools of pharmaceutical research to generate testable hypotheses about the most consumed bioactive beverage on Earth. Not to medicalize coffee. Not to put health claims on packaging. Not to replace the simple pleasure of the drink with anxiety about molecular targets. But to know. Because knowing is what scientists do, and because informed consumption is better than ignorant consumption, and because the questions are genuinely beautiful.
How does a roasted seed extract manage to interact coherently with dozens of biological targets across multiple organ systems? Why does this particular combination of compounds — forged by the Maillard reaction from precursors that evolved for entirely different purposes in the coffee plant — happen to fit so neatly into human receptor binding sites? Is this coincidence, evolutionary convergence, or something about the fundamental chemistry of life that we haven’t yet understood?
I don’t know. That’s the point of this chapter.
Closing the Laptop, Opening the Question
I want to end where I began — with the cup.
This book has been, for me, an exercise in applied curiosity. I am trained as a physicist. I work with the tools of pharmaceutical science. I wrote a book about coffee. The through-line is not the subject matter but the method: take something familiar, look at it with quantitative tools, and see what reveals itself.
What revealed itself was more complex, more organized, and more surprising than I expected. Coffee is not a simple stimulant delivery system. It is a multi-component bioactive mixture with predicted interactions across virtually every major physiological system. Our computational studies generated dozens of specific, testable hypotheses — some confirming what epidemiology had long suggested, others (like the NR4A1 connection) pointing in directions nobody had looked.
But I want to be honest about the feeling that dominates as I finish this project. It is not triumph. It is not the satisfaction of problems solved. It is the slightly vertiginous awareness — the feeling you get standing at the edge of something vast — of how much territory lies between where we are and where we need to be. I finished my last simulation run on a Thursday evening. I closed the laptop, looked at the cold espresso on my desk, and thought: I know less about you than when I started. That’s not a failure. That’s what real science feels like.
We know more than we did before these four studies. We know far less than we need to. That gap — between what computation predicts and what biology confirms — is where the next decade of coffee science lives.
I find that gap exciting rather than discouraging. Every unknown on the list I described above is a research question. Every unvalidated prediction is an experiment waiting to be designed. Every limitation of our models is an invitation to build better ones.
The computational tools will continue to improve. The experimental techniques will become more sensitive. The data will accumulate. And slowly, study by study, prediction by prediction, confirmation by confirmation, we will close the distance between the model and the molecule, between the simulation and the cell, between the screen and the cup.
I’ll be here, at my desk, running the simulations. The coffee will be beside me — a little cold, as usual, because I got distracted by the results. The screen will be glowing with binding energy curves and network graphs. And somewhere in those patterns, the next surprise will be waiting.
Come along. The cup is still full of questions.
Next: Chapter 15 — Your Cup, Your Science
Patterns in your own coffee preferences that reveal your personal chemistry
You'll Need
- A notebook or spreadsheet
- Your daily coffee routine
- 30 days of commitment
Do This
- Each day, record: origin/blend, brew method, grind size, water temp if known, and a 1-5 enjoyment rating.
- Add one line about how you felt 2 hours later (energy, focus, mood).
- Note any food eaten with or before coffee.
- At day 15, look for patterns. At day 30, analyze your data.
- Identify your top 3 combinations and your worst. You now have a personal dataset.
What's Happening
You're building an n=1 dataset that captures the intersection of everything in this book: compound chemistry (origin, roast), extraction physics (brew method, grind), pharmacokinetics (your personal caffeine metabolism), and chronobiology (time of day). No study can tell you YOUR optimal cup — but 30 data points from your own life can. This is citizen science at its most practical.
“Every cup you drink from here on carries a story — a billion molecular events between the tree and your brain. Now you know how to read it.”