Chapter 6: The Hub Proteins
Chapter 6: The Hub Proteins
Part II: Coffee’s Hidden Network
The barista’s name was Marta, and she ran a tiny specialty café in the Gracia neighborhood of Barcelona — the kind of place with exactly twelve seats, a single-group La Marzocca, and an encyclopedic knowledge of Ethiopian processing methods. I had been coming there for months, usually with my laptop, usually lost in spreadsheets of protein interaction data. One afternoon she caught me frowning at a particularly tangled network diagram and asked what I was working on.
“I’m trying to figure out which proteins matter most in this network,” I told her.
She looked at the screen — a web of nodes and edges that probably looked like a subway map designed by someone having a bad day — and said, “It looks like a mess.”
“It is a mess,” I admitted. “But it’s a structured mess. That’s the whole point.”
I tried to explain it to her in terms she’d recognize.
“Imagine your café is a network,” I said. “Every customer, every supplier, every barista, every piece of equipment — those are nodes. The connections between them are edges. A customer talks to you. You pull a shot on the machine. The machine needs beans from the supplier. Information and material flow through the network along these edges.”
She was following.
“Now here’s the question: which node in your café network is the most important?”
She didn’t hesitate. “Me. Obviously. Everything goes through me.”
“Exactly,” I said. “You’re what network scientists call a hub. You have more connections than any other node. You connect the customer to the coffee, the supplier to the cup, the equipment to the output. If I removed any single customer, the café keeps running. If I removed you—”
“Chaos,” she said.
“Chaos,” I agreed. “The whole system stops.”
I turned the laptop back toward me. “That’s what I’m looking for in this network. The biological equivalent of you. The proteins that connect everything, the ones where if you removed them, entire pathways would go silent.”
What I didn’t tell Marta that afternoon — because it would have required another hour and a second cortado — was that I had already found them. Three proteins, sitting at the center of coffee’s biological network, each controlling an entire domain of human physiology. And their names were Nrf2, COX-2, and PPARγ.
Hub Proteins — The Traffic Controllers
In any complex network — social, technological, biological — the connections are not evenly distributed. This is one of the most important discoveries in modern network science, and it applies as much to protein interaction networks as it does to the internet or airline route maps.
Most nodes in a network have relatively few connections. A handful of nodes have an enormous number. These highly connected nodes are called hubs, and they exert disproportionate control over how information flows through the system. In an airline network, hubs are places like Atlanta, Dubai, or Frankfurt — remove one of those airports, and thousands of routes collapse. In a protein network, hubs are the proteins that interact with many other proteins, controlling entire signaling cascades and metabolic pathways.
When I built the network pharmacology model for coffee’s bioactive compounds — mapping which compounds are predicted to interact with which protein targets, and how those targets connect to each other — the network that emerged had 6 compounds, 10 protein targets, and 36 edges linking them together. And like every real biological network I’ve ever analyzed, it was not random. It was scale-free: a small number of proteins had far more connections than the rest.
Three proteins stood out as clear hubs, each sitting at the center of a different biological domain. And the fact that multiple coffee compounds are predicted to interact with all three of them began to explain something that had puzzled researchers for years: why coffee’s epidemiological associations are so remarkably broad.
Let me introduce them one at a time.
Nrf2 (NFE2L2) — The Antioxidant Master Switch
If I could nominate one protein as the most important node in coffee’s biological network, it would be Nrf2 — formally known as nuclear factor erythroid 2-related factor 2, encoded by the gene NFE2L2. Nrf2 is a transcription factor, which means it doesn’t do the biochemical work itself. Instead, it controls the expression of other genes — hundreds of them — that collectively mount the cell’s defense against oxidative stress.
Here is the analogy I find most useful. Think of Nrf2 as a fire alarm system in a large building. The building has sprinklers in every room, fire doors on every floor, extinguishers in every hallway, and an emergency communication system that alerts all occupants simultaneously. Each of these defensive elements works independently once activated. But the thing that activates all of them at once — the master switch that coordinates the entire defense — is the fire alarm. That’s Nrf2.
Under normal conditions, Nrf2 is kept on a short leash. It’s bound to a regulatory protein called Keap1 in the cell’s cytoplasm, which tags Nrf2 for degradation — essentially sending it to the cellular recycling bin before it can do anything. This keeps the antioxidant defense system in standby mode. Energy is conserved. The sprinklers stay off.
But when the cell detects oxidative stress — an accumulation of reactive oxygen species, electrophilic chemicals, or other signals of damage — the Keap1-Nrf2 interaction is disrupted. Nrf2 breaks free, travels to the nucleus, and binds to a specific DNA sequence called the antioxidant response element (ARE). This activates the transcription of a whole battery of protective genes: genes for glutathione synthesis, genes for heme oxygenase, genes for NAD(P)H quinone oxidoreductase, genes for drug-metabolizing enzymes. The fire alarm has gone off, and every sprinkler in the building starts working simultaneously.
What makes Nrf2 so central to coffee’s network is that multiple coffee compounds are predicted to interact with this single protein. Our network analysis indicates that chlorogenic acid, cafestol, and ferulic acid all show predicted interactions with Nrf2. These are structurally diverse molecules — a polyphenol, a diterpene, and a hydroxycinnamic acid — arriving at the same master switch through different molecular pathways.
This is not redundancy. It’s convergence. And it has a practical implication: even if one compound is present at a low concentration, or is partially degraded by roasting, or is poorly absorbed in the gut, the other compounds may still reach Nrf2 and activate the same downstream defense. The system has built-in resilience, not because anyone designed it that way, but because that’s how hub-based networks naturally behave.
What this means for your cup: Because three structurally different compounds converge on Nrf2, your roast level and brewing method change which compounds reach the target — but not whether you hit it. A light roast delivers more CGA-derived activators. A dark roast delivers more cafestol. Both pathways lead to the same master switch. Your antioxidant defense gets activated either way — through different doors.
I remember the moment I realized this convergence pattern in my data. I was expecting each compound to interact with its own unique set of targets — a one-to-one mapping, neat and tidy. Instead, the network kept pointing back to Nrf2, over and over, like a subway map where every line passes through the same central station. It wasn’t what I expected. It was far more interesting.
If you think Nrf2 sounds important, the pharmaceutical industry agrees with you. As of the early 2020s, dozens of drugs in various stages of development target the Nrf2-Keap1 pathway. The logic is simple: if you can activate Nrf2, you turn on the cell’s own antioxidant defense system rather than trying to supply antioxidants from the outside (which, as decades of disappointing vitamin E trials have shown, doesn’t work very well).
The most advanced Nrf2-targeting drug is dimethyl fumarate (marketed as Tecfidera), approved for the treatment of multiple sclerosis. It works, at least in part, by activating Nrf2 and boosting the brain’s antioxidant capacity.
Now, I want to be absolutely clear: I am not saying that coffee is a substitute for pharmaceutical Nrf2 activators. The doses, the pharmacokinetics, the clinical contexts are entirely different. But the fact that coffee compounds are predicted to interact with the same master switch that pharmaceutical companies have spent billions targeting is, at the very least, a striking coincidence. And it suggests that the molecular basis of coffee’s epidemiological associations may not be as mysterious as it once seemed.
COX-2 (PTGS2) — The Inflammation Mediator
The second hub in coffee’s network is a protein you have almost certainly interacted with, even if you’ve never heard its name. COX-2 — cyclooxygenase-2, encoded by the gene PTGS2 — is the enzyme that produces prostaglandins, the chemical messengers that drive inflammation, pain, and fever. When you stub your toe and it swells and throbs, COX-2 is the molecular reason.
COX-2 is also the target of some of the most widely consumed drugs in human history. When you take aspirin, you are inhibiting COX-2 (and its cousin COX-1). When you take ibuprofen, naproxen, or celecoxib, same story. The entire class of non-steroidal anti-inflammatory drugs (NSAIDs) works primarily by blocking COX enzymes. This makes COX-2 one of the most pharmaceutically validated drug targets in existence — a protein whose biological importance has been confirmed by over a century of clinical medicine.
In coffee’s network, COX-2 appeared as a hub because multiple coffee compounds are predicted to interact with it. Our analysis indicates that these interactions likely occur through mechanisms distinct from how aspirin or ibuprofen bind to the enzyme. This is an important nuance. Aspirin irreversibly acetylates COX-2, physically modifying the enzyme’s structure. Ibuprofen competes for the enzyme’s active site, blocking the substrate from entering. The predicted interactions of coffee’s polyphenols with COX-2 appear to involve different binding regions and different molecular mechanisms — more like modulation than outright blockade.
This distinction matters because it may explain something about the safety profile of habitual coffee consumption. NSAIDs are effective anti-inflammatory agents, but chronic use carries well-known risks: gastrointestinal bleeding, kidney damage, cardiovascular complications at high doses. Coffee, consumed daily by billions of people for centuries, is associated with none of these specific problems at moderate intake levels. If coffee compounds are predicted to interact with COX-2 through gentler, modulatory mechanisms rather than the blunt inhibition of pharmaceutical NSAIDs, that could partially explain why.
Now, this is the point where I need to be particularly careful with language. Epidemiological studies have consistently found that regular coffee consumption is associated with lower levels of systemic inflammatory markers — C-reactive protein, interleukin-6, tumor necrosis factor alpha. These are associations, not proof of causation. But the fact that our network analysis independently identifies COX-2 as a hub for coffee compound interactions offers a potential molecular mechanism that is consistent with the epidemiological data. The network suggests an explanation. It doesn’t prove it.
What I find intellectually satisfying about the COX-2 hub is how neatly it bridges the molecular and the epidemiological. For decades, researchers noted the anti-inflammatory associations of coffee consumption and had no good mechanistic story to tell. The network provides one: multiple compounds, converging on a single well-validated inflammatory mediator, through predicted interactions that are distinct from pharmaceutical inhibition. It doesn’t close the case. But it opens a very specific line of investigation.
Here’s a fact that I find endlessly fascinating: aspirin and coffee compounds are predicted to interact with the same protein — COX-2 — through completely different mechanisms.
Aspirin is a small molecule (molecular weight just 180 daltons) that irreversibly acetylates a serine residue in COX-2’s active site. It’s a covalent modification — aspirin physically changes the enzyme’s structure, permanently disabling it. The cell has to make entirely new COX-2 molecules to restore function. This is why a single aspirin dose has effects that last for hours: the enzyme it hit is dead, and replacement takes time.
Coffee’s polyphenolic compounds, by contrast, are predicted to interact with COX-2 through non-covalent binding — hydrogen bonds, van der Waals forces, hydrophobic interactions. These are reversible. The compound binds, modulates the enzyme’s activity, and eventually dissociates. No permanent modification. No enzyme destruction.
Think of the difference this way: aspirin is a sledgehammer. Coffee compounds are more like a hand on someone’s shoulder — a gentle pressure that slows them down without breaking anything. Both affect the same target. Both may reduce inflammatory signaling. But the nature and intensity of the interaction are fundamentally different.
This is why I find network pharmacology so powerful. It reveals shared targets that you would never discover by studying coffee and aspirin in isolation. The network doesn’t care about the history of pharmaceutical development or the sociology of beverage consumption. It just maps the molecular connections. And sometimes those connections are surprising.
Two hubs down, one to go. If you’re sensing a pattern — compound hits protein, protein controls a cascade, cascade explains an epidemiological mystery — you’re right. That is the pattern. But the third hub is the one that surprised me most, because it connects coffee to the health crisis I see every time I open a medical journal.
PPARγ — The Metabolic Regulator
PPARγ — peroxisome proliferator-activated receptor gamma — is a nuclear receptor that regulates genes involved in lipid metabolism, glucose homeostasis, and adipocyte (fat cell) differentiation. When PPARγ is activated, it turns on genes that improve insulin sensitivity, promote fatty acid storage in adipose tissue rather than in the liver or muscle, and modulate the inflammatory environment of metabolic tissues.
If Nrf2 is the fire alarm and COX-2 is the inflammation mediator, then PPARγ is the metabolic thermostat — the protein that helps maintain the balance between fat storage and fat burning, between insulin sensitivity and insulin resistance.
The pharmaceutical relevance of PPARγ is well established. A class of diabetes drugs called thiazolidinediones — including pioglitazone and the now-withdrawn rosiglitazone — work specifically as PPARγ agonists. They activate this receptor to improve insulin sensitivity in patients with type 2 diabetes. These drugs are effective, though they carry side effects (weight gain, fluid retention, and in the case of rosiglitazone, possible cardiovascular risks) that have complicated their clinical use.
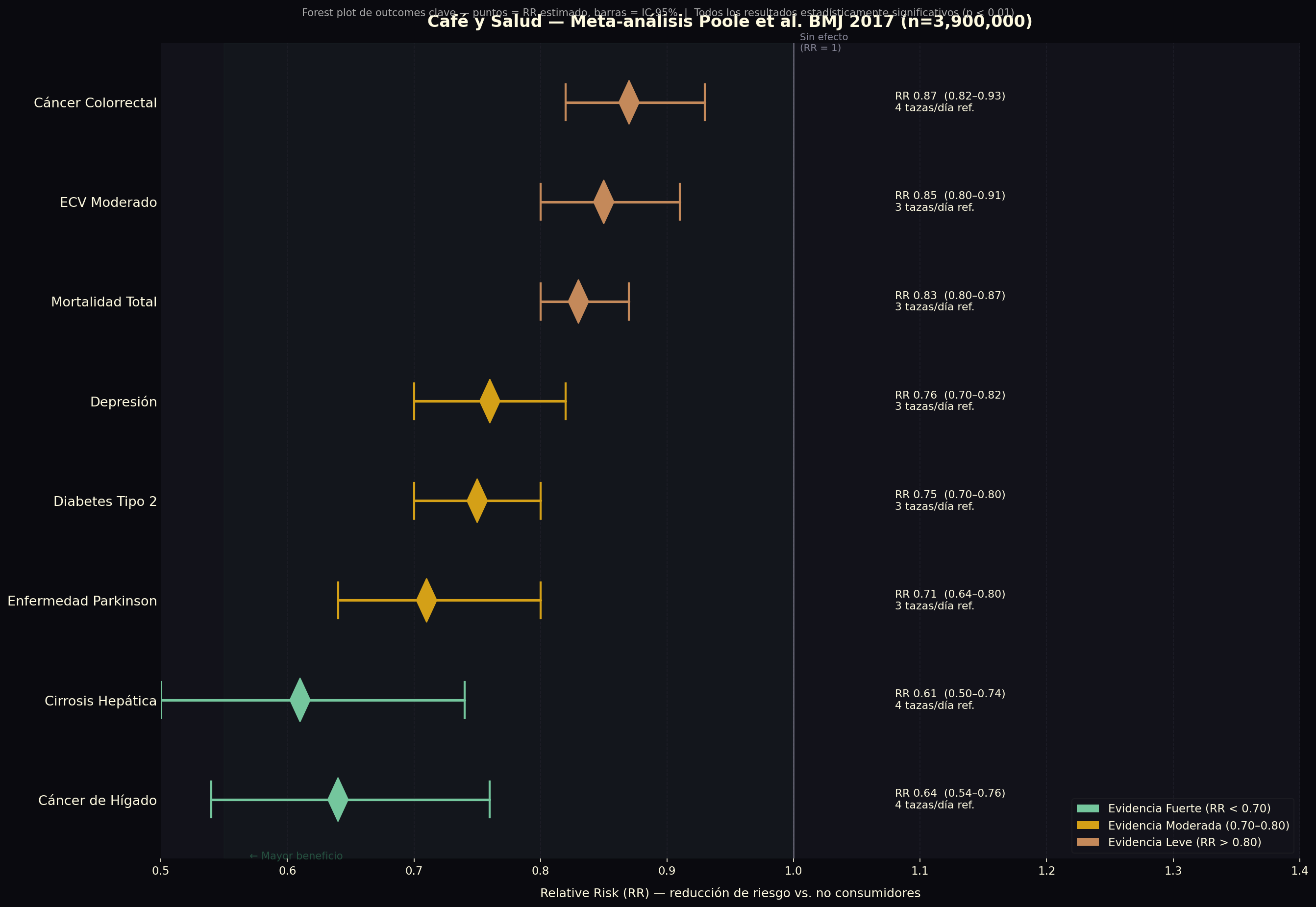
Our network analysis indicates that coffee compounds are predicted to interact with PPARγ. This prediction is particularly interesting in light of one of the most robust epidemiological findings in all of coffee research: the association between regular coffee consumption and reduced risk of type 2 diabetes. Meta-analyses consistently report an approximately 29% reduction in risk among habitual coffee drinkers compared to non-drinkers — one of the strongest and most replicated dietary associations in metabolic epidemiology.
For years, this association lacked a convincing molecular explanation. Caffeine alone doesn’t account for it — decaffeinated coffee shows similar associations. Chlorogenic acids have been proposed as candidates, based on their effects on glucose absorption in cell and animal studies. But the network offers something more specific: a convergence of multiple coffee compounds on PPARγ, a validated metabolic regulator, through predicted interactions that could plausibly modulate insulin sensitivity.
I want to be precise about what this means and what it doesn’t. The network analysis does not prove that coffee prevents diabetes. It does not establish that the PPARγ interactions are strong enough, sustained enough, or bioavailable enough to produce clinically meaningful effects. What it does is identify a specific, testable molecular hypothesis that is consistent with the epidemiological data. That’s how computational biology is supposed to work: not as proof, but as a generator of hypotheses worth testing.
What this means for you: If metabolic health is your concern, the epidemiology and the network analysis point the same direction — moderate, regular coffee consumption is associated with improved insulin sensitivity. The PPARγ hub offers a molecular explanation for why. And since decaf shows similar associations (remember, caffeine is not the only player), this effect appears to be driven by the polyphenols and other non-caffeine compounds. Your choice of decaf versus regular matters less here than whether you drink coffee at all.
NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) is a master transcription factor controlling inflammatory gene expression. Coffee polyphenols, particularly chlorogenic acid and its metabolites, are predicted to modulate NF-κB signaling by inhibiting IκB kinase (IKK) phosphorylation. This reduces the translocation of NF-κB to the nucleus, dampening the transcription of pro-inflammatory cytokines such as TNF-α, IL-6, and IL-1β. The convergence of multiple coffee compounds on this pathway provides a mechanistic link between habitual coffee consumption and the lower systemic inflammatory markers observed in epidemiological studies.
The Bridge Proteins — CYP1A2 and GSK3β
Hubs aren’t the only structurally important nodes in a network. There’s another category of protein that plays a different but equally critical role: bridge proteins.
In network science, a bridge is a node with high betweenness centrality — meaning that a large proportion of the shortest paths between other nodes pass through it. Hubs have many connections. Bridges may not have the most connections, but they sit at strategic bottlenecks between clusters. Remove a hub, and one cluster loses its organizing center. Remove a bridge, and two clusters that were connected become isolated from each other entirely.
In coffee’s biological network, two proteins emerged with the highest betweenness centrality: CYP1A2 and GSK3β. And each tells a fascinating story.
CYP1A2 — Your Caffeine Speed
CYP1A2 is a cytochrome P450 enzyme — one of the liver’s primary drug-metabolizing workhorses. Its day job is breaking down xenobiotics: foreign chemicals that enter the body through food, drink, drugs, and environmental exposure. And among those xenobiotics, one of CYP1A2’s most important substrates is caffeine.
When you drink a cup of coffee, caffeine is absorbed through the gastrointestinal tract, enters the bloodstream, crosses the blood-brain barrier, and begins blocking adenosine receptors within about 20 minutes. But it doesn’t stay in your system forever. CYP1A2, primarily in the liver, is responsible for approximately 95% of caffeine metabolism in humans. It converts caffeine into paraxanthine, theobromine, and theophylline — metabolites that are eventually excreted.
Here’s where it gets personal. The gene encoding CYP1A2 is polymorphic, meaning it comes in different versions in different people. The most well-studied variant is a single nucleotide change — a swap from A to C at a specific position — that determines whether you produce a fast or slow version of the enzyme.
If you carry the AA genotype, you’re a fast metabolizer. Your CYP1A2 enzyme chews through caffeine quickly, clearing it from your blood with a half-life of roughly three to four hours. You’re the person who can drink an espresso after dinner and sleep like a baby.
If you carry the AC or CC genotype, you’re a slow metabolizer. Your CYP1A2 works more gradually, and caffeine lingers in your system with a half-life that can stretch to six hours or more. You’re the person who can’t have coffee past 2 PM without staring at the ceiling at midnight.
Try this self-test: Drink a cup of coffee at 3 PM on a day you have nothing critical the next morning. Track how long it takes you to fall asleep. If you sleep normally, you’re likely a fast metabolizer. If you’re still wired at midnight, you’re likely slow. This is not a genetic test — but it’s a remarkably good proxy for your CYP1A2 phenotype, and it costs nothing.
In our network, CYP1A2 appeared as a bridge because it connects two otherwise separate cluster domains: the metabolic processing of coffee compounds and the downstream signaling effects of those compounds. It sits at the junction between “what happens to coffee in your body” and “what coffee does to your body.” Remove CYP1A2 from the network, and these two domains lose their primary connection.
This bridge position also explains why CYP1A2 genetic variation has such outsized effects on the coffee-health relationship. Studies suggest that the association between coffee consumption and cardiovascular outcomes may differ between fast and slow metabolizers — because the bridge protein that determines how quickly caffeine is cleared also determines how long caffeine-related signaling persists. The bridge doesn’t just connect; it modulates the flow between clusters.
You know that friend who drinks a double espresso at 9 PM and falls asleep by 10? And you know how you tried that once and were still wide awake at 2 AM? The difference is probably not willpower, habit, or tolerance. It’s genetics. Specifically, it’s your CYP1A2 gene.
About 40-45% of the population carries the fast-metabolizer genotype (AA). These people clear caffeine from their bloodstream relatively quickly — half of it gone in about three to four hours. For them, an afternoon coffee is ancient metabolic history by bedtime.
The remaining 55-60% carry one or two copies of the slow variant (AC or CC). Their CYP1A2 enzyme works at a more leisurely pace, and caffeine sticks around longer. A coffee at 3 PM can still have meaningful concentrations of caffeine circulating at midnight.
What’s remarkable is that this same genetic variation appears to influence the health associations of coffee consumption. Some research suggests that for fast metabolizers, moderate coffee intake is associated with reduced cardiovascular risk — perhaps because the beneficial non-caffeine compounds are absorbed while caffeine is cleared quickly. For slow metabolizers, the association is less clear, and some studies suggest it may even reverse at high intake levels — perhaps because prolonged caffeine exposure in slow metabolizers produces different hemodynamic effects.
You can find out your CYP1A2 status through consumer genetic testing services, though I’d caution against making dramatic lifestyle changes based on a single gene variant. Your response to coffee is influenced by dozens of genes, your microbiome, your habitual intake, your sleep architecture, and a hundred other variables. But if you’ve always wondered why coffee affects you differently than it affects your partner or your colleagues — CYP1A2 is very likely part of the answer.
GSK3β — The Bridge Between Brain and Body
The second bridge protein in coffee’s network is GSK3β — glycogen synthase kinase 3 beta. If CYP1A2 bridges metabolism and signaling, GSK3β bridges two of the most important pathological processes in modern medicine: neuroinflammation and oxidative stress.
GSK3β is a kinase — an enzyme that adds phosphate groups to other proteins, modifying their activity. It was originally discovered in the context of glycogen metabolism (hence the name), but research over the past two decades has revealed it to be a remarkably versatile signaling molecule involved in an extraordinary range of cellular processes: cell proliferation, apoptosis, gene expression, neuronal development, and immune response.
What makes GSK3β particularly interesting in the context of coffee is its dual role. On one side of the network, GSK3β connects to neuroinflammatory pathways — the chronic, low-grade inflammation in brain tissue that research has linked to neurodegenerative conditions like Alzheimer’s and Parkinson’s disease. On the other side, it connects to oxidative stress pathways — the Nrf2-mediated defense systems we discussed earlier in this chapter.
GSK3β is, in a sense, the protein that decides whether oxidative damage in the brain triggers an inflammatory cascade or an antioxidant defense response. It sits at the decision point between escalation and resolution. And our network analysis indicates that coffee compounds are predicted to interact with this critical bridge.
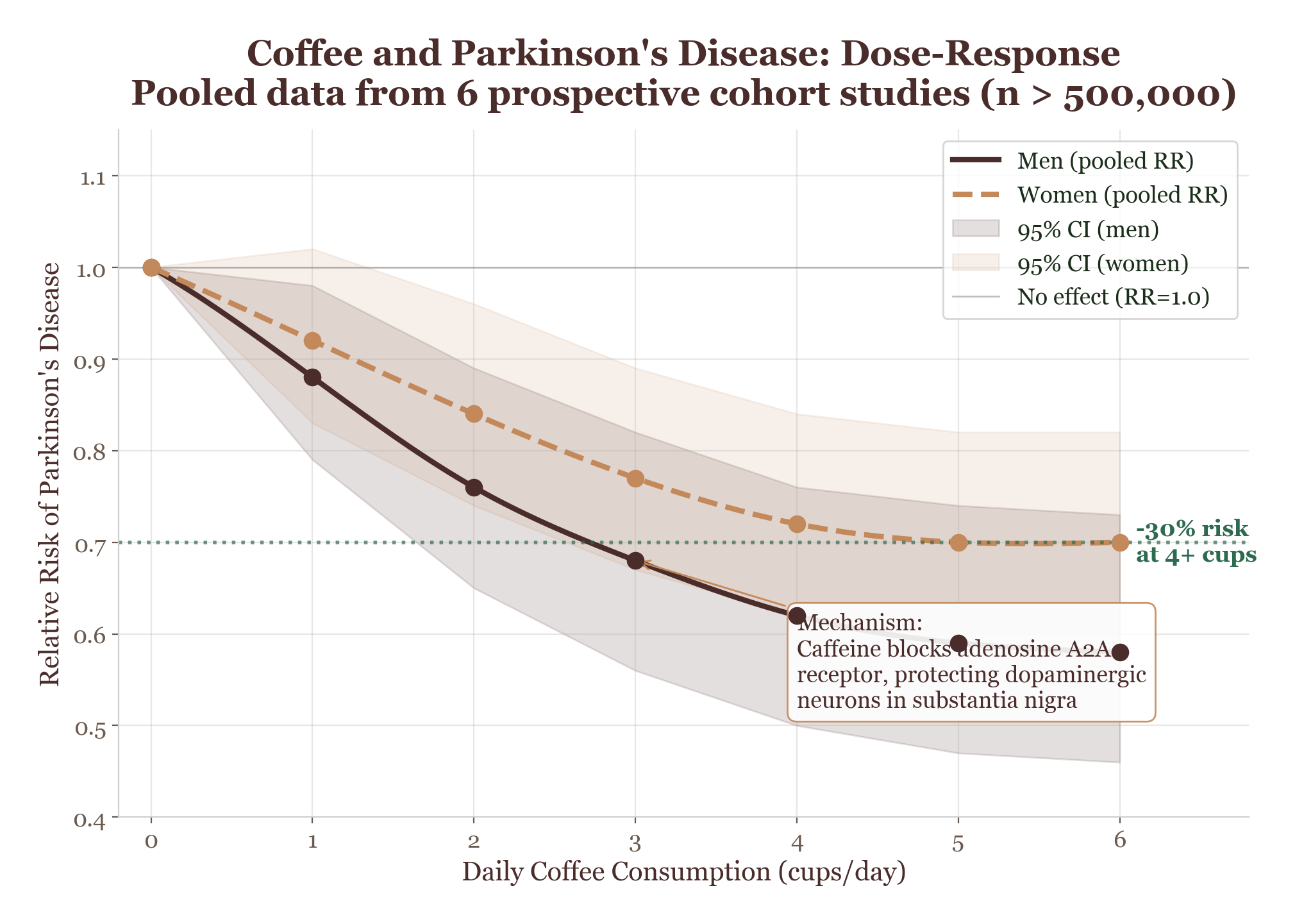
This bridge position may be relevant to one of the most intriguing epidemiological observations about coffee: the consistent association between regular coffee consumption and reduced risk of Parkinson’s disease (approximately 28% in meta-analyses) and, to a lesser extent, Alzheimer’s disease. These are neurodegenerative conditions where both neuroinflammation and oxidative stress play well-documented roles. A bridge protein that connects these two pathological processes, and that coffee compounds are predicted to interact with, offers a structural molecular explanation for why coffee’s neurological associations are so robust.
Again, I must emphasize: this is a hypothesis generated by network analysis, not a proof. The network indicates that GSK3β sits at a critical junction. The epidemiology indicates that coffee consumption is associated with neuroprotection. The computational model predicts that coffee compounds interact with this junction. These three observations are consistent with each other, which is promising, but consistency is not causation. Proving this mechanism would require clinical studies that, as of this writing, have not been conducted.
Why Hubs and Bridges Matter
Let me step back from the individual proteins for a moment and ask a bigger question: what does this network architecture mean for how we understand coffee?
The traditional way of thinking about dietary bioactives is reductionist. You isolate a compound — let’s say chlorogenic acid — and you test it against a battery of targets. You find that it interacts with this enzyme, modulates that receptor, scavenges this free radical. You compile a list of individual effects, and you try to add them up.
The problem with this approach is that it misses the structure. It treats all targets as equally important, all interactions as independent, all effects as additive. The network perspective reveals something fundamentally different.
Coffee’s biological network is organized around a small number of highly connected hub proteins — Nrf2, COX-2, PPARγ — each of which controls an entire domain of physiology. Multiple compounds converge on these hubs, creating a pattern of redundant targeting that makes the system robust to variation. If chlorogenic acid is degraded during roasting, ferulic acid may still reach Nrf2. If one compound is poorly absorbed, another may compensate.
The network is also organized around bridge proteins — CYP1A2, GSK3β — that connect otherwise separate pathway clusters. These bridges explain why coffee’s effects seem to span so many different domains of health: antioxidant defense, inflammation, metabolism, neuroprotection. The domains aren’t separate. They’re linked through shared bridge proteins, and coffee compounds are predicted to interact with those bridges.
This architecture has a name in network science: a small-world network. Picture it this way. You walk into a crowded party knowing no one. But the host — a hub — knows half the room. Introduce yourself to her, and suddenly you’re two handshakes from anyone. That’s what Nrf2, COX-2, and PPARγ do for coffee’s compounds. They’re the hosts at the molecular party. Any compound can reach nearly any biological process through one of these three proteins. It’s the same architecture found in airline routes, the internet, and the “six degrees of separation” phenomenon. Maximum connectivity, minimum wiring.
And it means that coffee’s biological effects are not random. They’re not the result of a thousand compounds hitting a thousand targets in a thousand uncoordinated ways. They’re the result of a focused set of compounds interacting with a small number of structurally critical proteins that each cascade across broad swaths of human physiology.
It’s elegant, not chaotic. And that elegance — the fact that a random plant extract happens to be organized like a well-designed drug — is the thing I still find hardest to explain and most thrilling to discover.
Here’s what I want you to take away from this chapter, the next time you’re holding your morning coffee.
That cup does not contain a random assortment of molecules doing random things to random targets in your body. Our network analysis indicates that it contains compounds predicted to interact with a small number of highly connected hub proteins — Nrf2, COX-2, PPARγ — each of which sits at the center of an entire physiological domain. Antioxidant defense. Inflammation. Metabolism. The compounds don’t need to hit a hundred targets to have broad effects. They hit a few hubs, and the hubs do the rest.
The network also contains bridge proteins — CYP1A2, GSK3β — that connect these domains to each other. Your genetic variant of CYP1A2 influences how quickly you process caffeine, which may shape how coffee affects you personally. GSK3β connects oxidative stress to neuroinflammation, which may help explain coffee’s associations with neurodegenerative disease.
None of this constitutes medical advice. I’m not telling you to drink more coffee or less coffee or different coffee. What I’m telling you is that coffee’s remarkably broad epidemiological profile — the associations with reduced risk of type 2 diabetes, Parkinson’s disease, certain liver conditions, and chronic inflammation — may have a structural molecular explanation. The network isn’t random. It’s organized around hubs and bridges. And coffee’s compounds appear to have found those hubs.
Whether they found them through millions of years of plant-animal co-evolution, or through sheer chemical coincidence, is a question I don’t yet know how to answer. But I find the architecture beautiful either way.
In the next chapter, we’ll zoom in even closer — to the atomic level — and watch what happens when one of these compounds actually slides into the binding pocket of a hub protein. If this chapter was the satellite view of the city, the next chapter is walking through the front door of a building. You’ll see atoms. You’ll see hydrogen bonds forming and breaking. You’ll see why shape matters more than chemistry.
Marta, back in her café in Sant Antoni, doesn’t know about Nrf2 or CYP1A2. She just knows that the cortado she’s pouring right now — the one with the crema settling like a copper penny — will reach five hub proteins before she wipes down the counter. I think she’d find that worth knowing.
How different brewing cultures accidentally optimize different chemical extractions
You'll Need
- Same coffee beans (medium roast)
- Turkish ibrik/cezve or small pot
- Pour-over setup
- Jar for cold brew
- Grinder
- Timer
Do This
- Turkish: Grind ultra-fine, boil 10g in 100ml water with 1tsp sugar, bring to foam three times.
- Pour-over: Grind medium, brew 15g with 240ml water at 93°C.
- Cold brew: Grind coarse, steep 30g in 300ml cold water for 12–18 hours (start this first!).
- Taste all three. Note strength, acidity, bitterness, sweetness.
What's Happening
Turkish coffee extracts maximum compounds (fine grind + boiling = high extraction). Pour-over extracts selectively (paper filter removes oils, controlled temperature). Cold brew extracts slowly (low temperature suppresses bitter compounds but extracts sugars and smooth oils). Three cultures, three centuries of tradition — each accidentally optimizing a different chemical equation.
“When you drink matters almost as much as what’s in it. Your body runs on a chemical clock — and coffee either works with it, or against it.”