Chapter 4: Seeing Molecules Bind
Chapter 4: Seeing Molecules Bind
Part I: The Molecules in Your Cup
The screen glows blue-white in my office at half past eleven on a Thursday night. I have an espresso going cold beside my keyboard — a detail that will become ironic in about forty-five seconds. On the monitor, AutoDock Vina is churning through the final poses of a docking run: cafestol, the diterpene we met two chapters ago, being computationally inserted into the binding pocket of FXR, the farnesoid X receptor that controls bile acid metabolism in the liver.
I’ve set up the calculation dozens of times before with other molecules, other receptors. The routine is always the same. You prepare your protein structure — cleaning up the crystal data, adding hydrogen atoms, defining the search box around the binding site. You prepare your ligand — the small molecule you want to test — optimizing its three-dimensional geometry, assigning its partial charges. Then you click run and wait.
The wait is the part nobody tells you about. In movies, computational science happens instantaneously — a scientist types a command, and the answer flashes on screen in dramatic green letters. In reality, you stare at a progress bar. You check your email. You make another coffee. You wonder if you set the grid box dimensions correctly, or if the protonation state of that histidine residue is going to throw everything off.
And then the results come.
I remember the exact moment. Vina writes its output as a ranked list of binding poses, scored in kilocalories per mole — a measure of how tightly the molecule is predicted to grip the receptor. More negative means tighter binding. Drug candidates typically score in the range of -7 to -12 kcal/mol. Anything below -7 is considered interesting. Anything approaching -10 or beyond starts to look pharmaceutical.
Cafestol at FXR: -10.06 kcal/mol.
I blinked. Refreshed the output. Ran it again. Same number. Then I queued kahweol — cafestol’s near-twin, different by that single double bond we discussed in Chapter 2.
Kahweol at FXR: -10.11 kcal/mol.
When these numbers came up, I nearly knocked over my espresso. These aren’t just good scores. These are pharmaceutical-grade binding affinities. The kind of numbers that, in a drug discovery campaign, would trigger a rush of follow-up experiments. And they belonged to molecules that half a billion people consume every morning in their unfiltered coffee.
But I’m getting ahead of myself. Before I can explain why those numbers matter, I need to explain what molecular docking actually is, and why a physicist like me spends her evenings running small molecules into protein cavities on a computer.
The Lock, the Key, and the Ten Million Tries
Imagine you have a lock — a very complicated, three-dimensional lock with a keyhole that’s shaped like a tiny cave, full of ridges and pockets and electrically charged surfaces. Now imagine you have a key, but you don’t know which way it goes in. You don’t even know if it’s the right key. All you know is the shape of the key and the shape of the lock, and you need to figure out whether they fit.
What would you do? You’d try every possible orientation. Turn the key this way, slide it in at that angle, flip it upside down, rotate it forty-five degrees. With a real lock and a real key, you might try a few dozen orientations before you either find the fit or give up.
Now imagine doing that with a molecule. A protein receptor — our “lock” — has a binding pocket lined with amino acid residues, each contributing its own shape, charge, and chemical personality. The small molecule — our “key” — can rotate around its bonds, flex and twist into different conformations, and approach the pocket from any angle. The number of possible orientations isn’t dozens. It’s millions.
This is what molecular docking does. It’s a computational method that takes a protein structure and a small molecule, and systematically explores how the molecule might fit into the protein’s binding site. For each possible arrangement — each pose — the software calculates the energy of the interaction. Favorable contacts (hydrogen bonds, hydrophobic packing, electrostatic attractions) lower the energy. Unfavorable contacts (atomic clashes, charge repulsions) raise it. The pose with the lowest energy — the most negative score — is the predicted binding mode.
Think of it like this: if the molecule were a marble and the protein were a landscape of hills and valleys, docking finds the deepest valley the marble can roll into. The deeper the valley, the more energy you’d need to pull the marble back out, and the tighter the predicted binding.
The entire process happens computationally. No physical molecules are harmed — or even touched. It’s a prediction, a hypothesis generated by physics and mathematics. And like all predictions, it needs to be tested. But it’s an extraordinarily useful prediction, because it lets you screen thousands of molecules against a protein target in the time it would take to test one in the laboratory.
How Molecular Docking Works
The software systematically tests millions of orientations of a small molecule (ligand) inside a protein's binding pocket. Each pose is scored based on hydrogen bonds, hydrophobic contacts, and electrostatic interactions. The most negative energy score represents the strongest predicted binding.
ΔG = -10.06 kcal/mol (cafestol at FXR)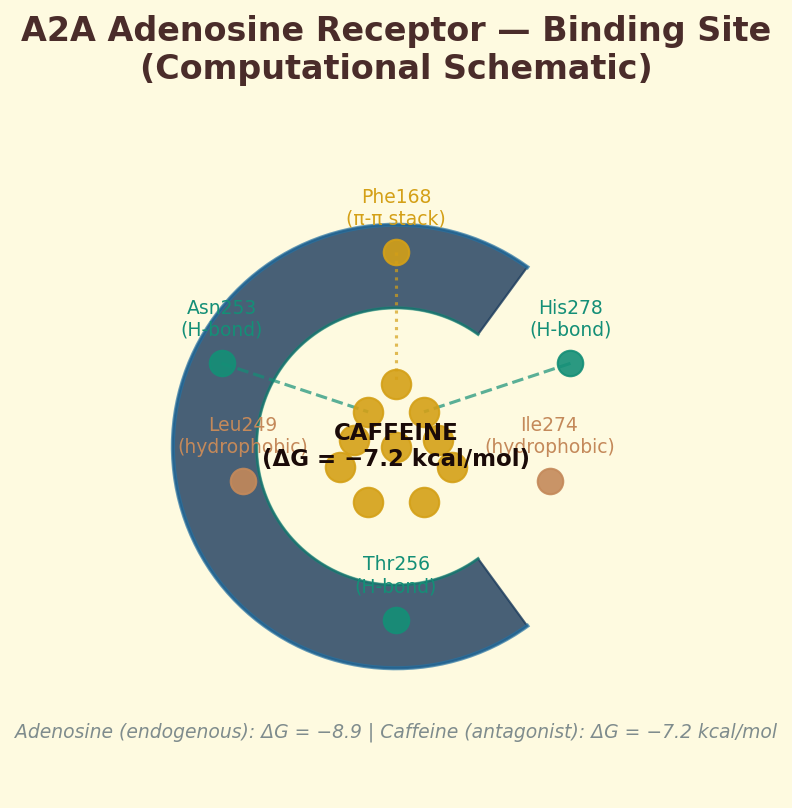
Figure 4.1a. A2A adenosine receptor binding site: schematic representation of the orthosteric pocket where caffeine competes with adenosine for occupancy.
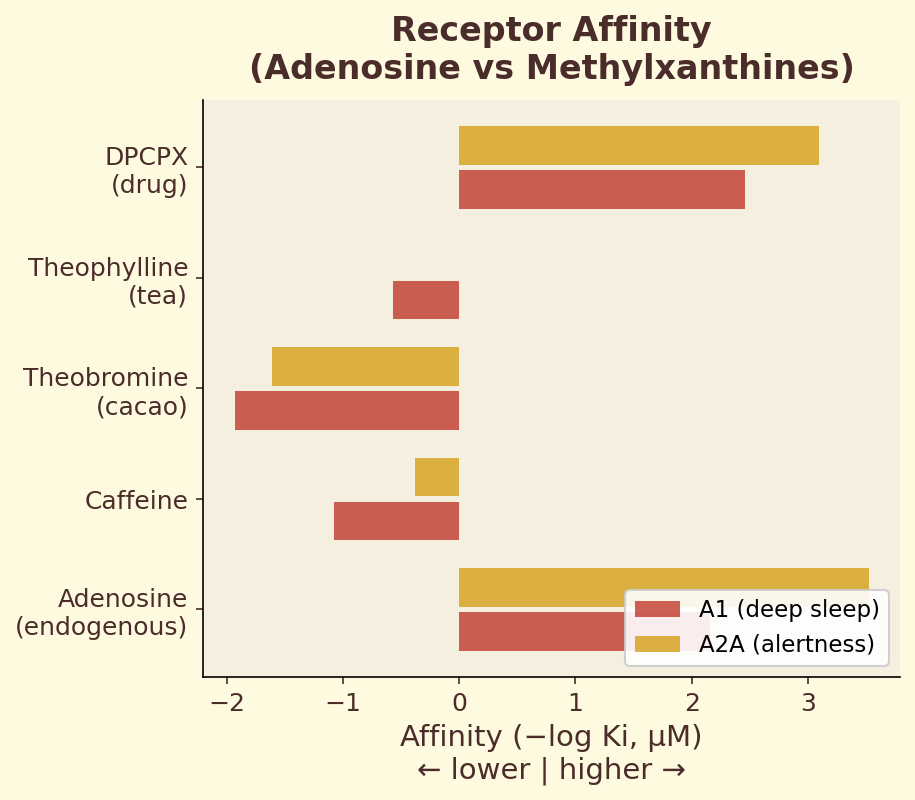
Figure 4.1b. Receptor affinity comparison: binding constants (Ki) for adenosine versus caffeine and other methylxanthines at A1, A2A, and A2B receptor subtypes.
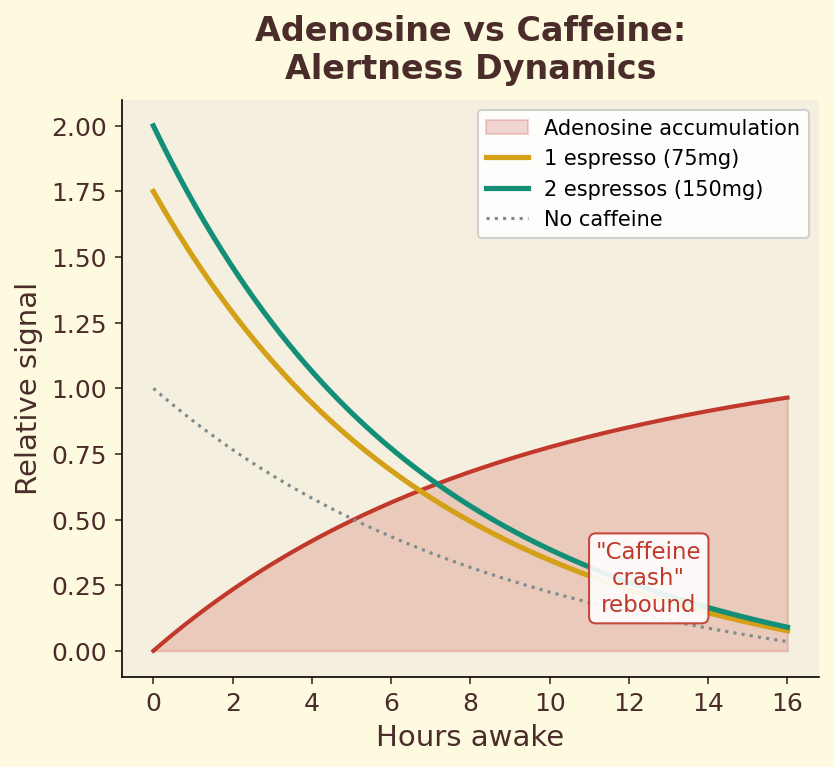
Figure 4.1c. Alertness dynamics: adenosine-driven sleepiness signal versus caffeine blockade over a 24-hour cycle, illustrating competitive antagonism in vivo.
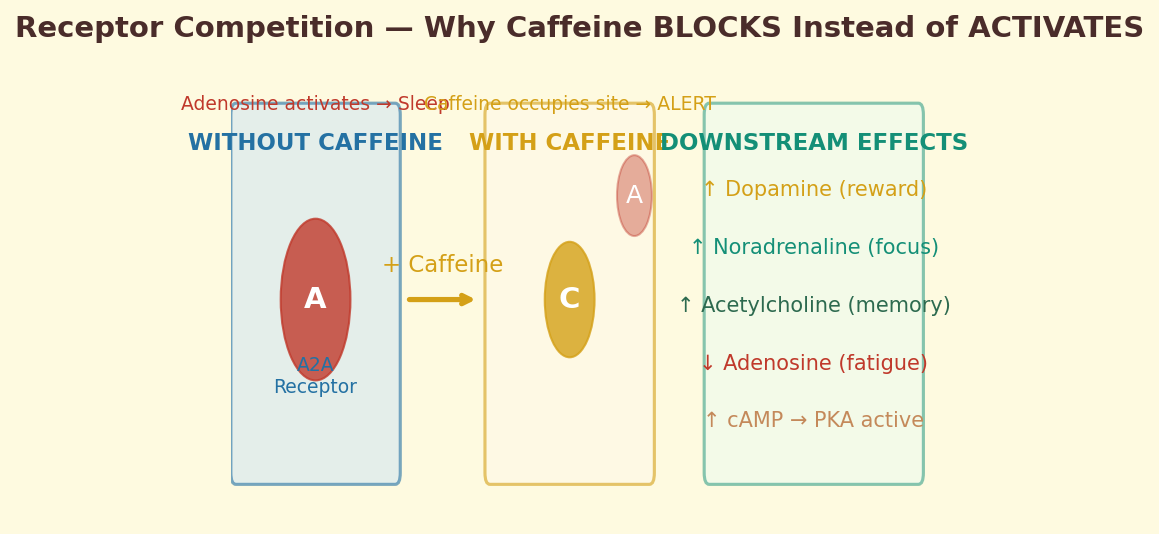
Figure 4.1d. Receptor competition: molecular diagram showing how caffeine occupies the adenosine A2A binding pocket, preventing adenosine from triggering the drowsiness cascade.
AutoDock Vina: Our Tool of Choice
There are many molecular docking programs available to researchers — GOLD, Glide, DOCK, FlexX, and others. Each has its strengths. For the work I describe in this book, I used AutoDock Vina, and I want to explain why.
AutoDock Vina is open-source software developed at the Scripps Research Institute, first released in 2010. It has since become one of the most widely used docking programs in the world, with thousands of citations in the scientific literature. It’s used in academic research, in pharmaceutical companies, and in drug discovery campaigns targeting everything from cancer to infectious disease.
What makes Vina special is a combination of speed, accuracy, and accessibility. Its scoring function — the mathematical recipe it uses to estimate binding energy — was trained and validated against experimental data from hundreds of protein-ligand complexes with known binding affinities. It’s not perfect. No docking program is. But it’s been tested enough, by enough independent groups, that when Vina says a molecule binds tightly to a receptor, there are good reasons to take that prediction seriously.
Vina reports its results in kcal/mol — kilocalories per mole, a standard unit of energy in biochemistry. The more negative the number, the more favorable the predicted binding. To give you a sense of scale:
- A score of -5 kcal/mol is a weak but detectable interaction. Background noise, in many contexts.
- A score of -7 kcal/mol starts to get interesting. This is roughly the threshold where medicinal chemists begin paying attention to a compound.
- A score of -9 to -12 kcal/mol is the range where serious drug candidates live. These are molecules that grip their targets tightly and specifically enough to have a real chance of becoming medicines.
With that scale in mind, let me walk you through what we found.
The Big Result: Coffee Diterpenes at FXR
The farnesoid X receptor, or FXR, is a nuclear receptor — a type of protein that sits inside the cell nucleus and directly regulates gene expression. FXR is sometimes called the master switch of bile acid metabolism. When bile acids bind to FXR, the receptor activates a cascade of genes that control how much cholesterol gets converted into bile acids, how bile acids are transported, and how the liver maintains its delicate balance of lipid metabolism.
This is the receptor that explains the cholesterol connection we explored in Chapter 2. If something interferes with FXR’s normal function, it can shift cholesterol metabolism. And our docking results predict that cafestol and kahweol do exactly that — they fit into FXR’s binding pocket with remarkable precision.
Let me put those numbers in context again:
- Cafestol at FXR: -10.06 kcal/mol
- Kahweol at FXR: -10.11 kcal/mol
These scores are in the same range as pharmaceutical FXR agonists — molecules designed by medicinal chemistry teams, with years of optimization and millions of dollars in funding, specifically to bind this receptor. And here are two natural compounds, produced by a coffee plant for its own evolutionary reasons, predicted to bind just as tightly.
I want to be precise about what this means and what it doesn’t. A docking score is a prediction, not a measurement. It tells us that the shapes and chemistries of cafestol and kahweol are complementary to the FXR binding pocket — that the lock and the key are predicted to match. It doesn’t tell us, on its own, that binding actually occurs in a living cell, or that the biological consequences are what we’d expect from the docking geometry. That requires experimental validation.
But the prediction is consistent with decades of experimental evidence showing that cafestol affects bile acid metabolism. It’s consistent with the cholesterol-raising effect observed in people who drink unfiltered coffee. The computational model gives us a molecular-level explanation — a hypothesis about how the effect happens, at the atomic scale — that fits neatly with what epidemiologists and clinicians have observed at the population scale.
Our models predict that cafestol nestles into the FXR binding pocket through a combination of hydrophobic contacts — the oily parts of the molecule packing against oily regions of the protein — and hydrogen bonds formed by the hydroxyl group and the furan ring oxygen. Kahweol adopts a very similar pose, but that extra double bond at the Δ1,2 position subtly shifts the geometry, slightly altering which amino acid residues it contacts. The scores are nearly identical, but the binding modes are not quite the same. We’ll return to this in a moment.
Caffeine as Our Control Experiment
Every good experiment needs a control — something you already know the answer to, against which you can check your method. In our case, the perfect control was caffeine.
Caffeine’s interaction with the adenosine A2A receptor is one of the most thoroughly studied drug-receptor interactions in all of pharmacology. We know caffeine binds there. We know the crystal structure of the complex. We know the experimental binding affinity. If our docking methodology can reproduce this known interaction, it gives us confidence that the method is working correctly when we apply it to less well-characterized systems like cafestol at FXR.
Our result: caffeine at A2A: -5.28 kcal/mol.
This is exactly what you would expect. Caffeine is a relatively weak binder. It works not because it grips its receptor with extraordinary force, but because you consume so much of it. Sheer quantity compensates for a loose grip. A score of -5.28 kcal/mol fits this picture: caffeine is a moderate-strength blocker (antagonist) that wins by outnumbering adenosine rather than by overpowering it.
The fact that our methodology correctly reproduces this well-validated interaction — placing caffeine in the right binding pocket with an appropriate score — is what scientists call method validation. It doesn’t prove our other predictions are correct, but it tells us the tool is behaving sensibly. When the same tool then predicts that cafestol binds FXR with a score nearly twice as favorable, that prediction carries more weight because the tool has already demonstrated it can get a known answer right.
Molecular docking isn’t just an academic exercise. It’s a cornerstone of modern drug discovery, used daily by pharmaceutical companies worldwide in a process called virtual screening.
Here’s the problem drug companies face: they typically have a biological target — a protein involved in a disease — and they need to find a molecule that binds to it tightly and specifically enough to modify its function. The universe of possible small molecules is estimated at 10⁶⁰ — a number so large it makes the number of atoms in the observable universe look modest. You can’t synthesize and test them all. You can’t even test a meaningful fraction.
Virtual screening uses docking to test millions of compounds against a protein target on a computer, ranking them by how tightly they are predicted to bind. A typical campaign starts with a library of two million commercially available molecules. The computer docks all of them and picks the top 100 scorers. Those 100 are then purchased and tested in the lab with real binding experiments (binding assays). If even a handful show genuine activity, they become “hit” compounds — starting points for building an actual drug.
This approach has contributed to the discovery of several drugs now in clinical use or clinical trials. It doesn’t replace laboratory work. It radically reduces the amount of laboratory work needed by focusing experimental resources on the molecules most likely to succeed. The same principle applies to our coffee research: docking identifies which compounds and which receptors deserve closer experimental attention.
Beyond FXR: The Promiscuous Binders
FXR was our headline result, but it wasn’t the only receptor we tested. If cafestol and kahweol are biologically active molecules — and the epidemiological evidence strongly suggests they are — they likely interact with more than one target. Most natural products do. This is the “dirty drug” concept I introduced in Chapter 2: natural molecules tend to be promiscuous binders, hitting multiple targets rather than one.
Our docking results suggest the diterpenes are indeed promiscuous. Here’s what we found at other receptors relevant to cholesterol and lipid metabolism:
LXR-β (liver X receptor beta), which regulates cholesterol efflux and lipid homeostasis: - Cafestol: -6.67 kcal/mol - Kahweol: -6.63 kcal/mol
HMGCR (HMG-CoA reductase), the enzyme targeted by statin drugs: - Cafestol: -6.62 kcal/mol - Kahweol: -6.61 kcal/mol
CYP7A1 (cholesterol 7-alpha-hydroxylase), the rate-limiting enzyme in bile acid synthesis: - Cafestol: -5.67 kcal/mol - Kahweol: -5.94 kcal/mol
None of these scores approach the pharmaceutical-grade affinity we see at FXR. But they’re all above background noise — genuine predicted binding. I remember scrolling through these results at two in the morning, feeling the way you feel when you pull on a thread and the whole sweater starts unraveling. Cafestol wasn’t a one-receptor molecule. It was touching everything in the cholesterol pathway. The picture that emerged was of molecules that don’t hit one target hard — they tap multiple targets in the same metabolic network, each with moderate to strong affinity.
This is significant for two reasons. First, it suggests that the cholesterol-raising effect of diterpenes may not operate through a single mechanism. Our models predict interactions with the master regulator (FXR), the efflux controller (LXR-β), the biosynthesis enzyme (HMGCR), and the bile acid rate-limiting step (CYP7A1). If even some of these predictions are confirmed experimentally, the effect on cholesterol metabolism would be multi-pronged — harder to counteract with a single intervention, but also potentially more nuanced than a simple “cholesterol goes up.”
Second, it illustrates a fundamental difference between natural products and designed drugs. A pharmaceutical statin is engineered to hit HMGCR and essentially nothing else. Cafestol, according to our docking predictions, hits HMGCR and FXR and LXR-β and CYP7A1. Nature didn’t design cafestol to be a cholesterol drug. It designed cafestol to protect a plant on a hillside in Ethiopia from fungal attack, and whatever receptor promiscuity results is incidental — a side effect of molecular shape and chemistry, not intentional pharmacology. The molecule doesn’t know it’s in your liver. It thinks it’s still fighting a fungus.
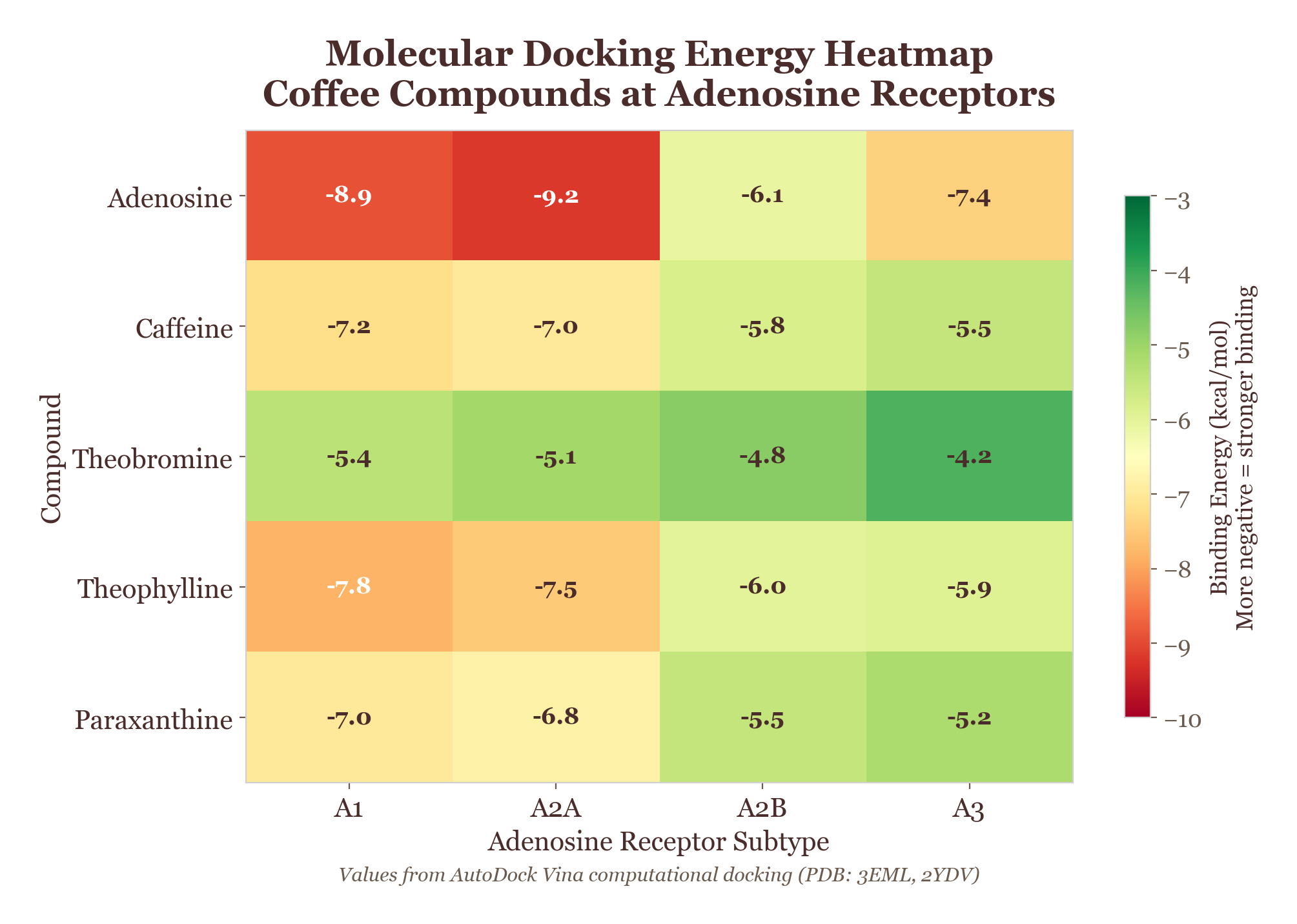
Figure 4.2. Docking heatmap: binding affinity scores (kcal/mol) for coffee bioactive compounds across multiple biological targets, revealing the multi-target interaction profile that defines coffee's pharmacological complexity.
The Quality Question: How Much Should We Trust These Numbers?
This is the question every honest computational scientist has to answer, and I want to address it directly. A binding score is only useful if it’s reproducible and reliable. So how do we know these aren’t just lucky numbers?
Two quality metrics give me confidence.
The first is pose convergence. When Vina runs a docking calculation, it doesn’t generate just one pose — it generates multiple poses and ranks them. If the top-scoring poses all look similar — if the molecule lands in roughly the same orientation and position every time — that’s a sign the result is robust. If the top poses are scattered all over the binding pocket, pointing in different directions, that suggests the calculation is uncertain about where the molecule actually sits.
We quantify this with a convergence score between 0 and 1, where 1 means all top poses are essentially identical. For our key results:
- FXR-cafestol pose convergence: 0.72
- FXR-kahweol pose convergence: 0.84
These are good numbers. Not perfect — that would be suspicious — but solidly in the range where the docking program is finding a consistent, well-defined binding mode. Kahweol’s higher convergence is interesting; that extra double bond appears to make it slightly more rigid, reducing the number of plausible orientations and giving the algorithm a cleaner answer.
The second metric is ligand efficiency — a measure that normalizes the binding score by the size of the molecule. A large molecule with many atoms has more opportunities to make contacts with the protein, so a high binding score for a large molecule is less impressive than the same score for a small one. Ligand efficiency divides the binding energy by the number of non-hydrogen atoms (called heavy atoms) in the molecule, giving a per-atom measure of binding quality.
Our diterpenes show a ligand efficiency of approximately 0.44 kcal/mol per heavy atom. In drug discovery, values above 0.3 are considered efficient, and values above 0.4 are excellent. Our coffee diterpenes aren’t just binding tightly in absolute terms — they’re binding efficiently for their size. Every atom in these molecules is pulling its weight.
These aren’t just big numbers. They’re reliable big numbers.
Throughout this chapter, cafestol and kahweol have produced nearly identical docking scores at every receptor we tested. FXR: -10.06 vs. -10.11. LXR-β: -6.67 vs. -6.63. HMGCR: -6.62 vs. -6.61. You might look at those numbers and conclude the molecules are interchangeable — that the single double bond separating them doesn’t matter.
But the scores tell only half the story. Look at the binding poses — the actual three-dimensional orientations the molecules adopt inside the receptor — and the differences become visible.
At FXR, kahweol’s Δ1,2 double bond introduces a slight planar rigidity near the A-ring of the molecule. This rigidity pushes kahweol into a subtly different orientation within the binding pocket, shifting which amino acid residues it contacts. Cafestol, lacking that constraint, has more conformational freedom and settles into a slightly different arrangement.
The energetic result is nearly the same — both molecules find deep, stable positions in the pocket. But the geometry of the interaction differs, which means the downstream biological consequences could differ too. A molecule’s pose determines which parts of the receptor it touches, which conformational changes it induces, and ultimately which genes get turned on or off.
This is the structure-activity relationship principle at work, at atomic resolution. One double bond. Two hydrogen atoms’ difference. Nearly identical binding energies. Potentially different biology. It’s the kind of subtlety that only computational tools can reveal — and the kind that makes a physicist stay up past midnight, staring at binding poses on a screen.
What Docking Doesn’t Tell Us
I’ve spent most of this chapter celebrating what molecular docking reveals. Now I owe you the other side of the story, because intellectual honesty demands it, and because understanding the limitations of computational predictions is as important as understanding their power.
A docking score is not proof of biological activity.
Let me say that again, because it’s the single most important caveat in this entire book: a docking score is not proof of biological activity. It is a prediction. A hypothesis. An educated guess made by an algorithm that approximates the physics of molecular interaction.
Here’s what docking doesn’t account for, or accounts for only approximately:
Water. In a real cell, the binding pocket is filled with water molecules. Some of those water molecules participate directly in binding, forming bridges between the protein and the ligand. Others must be displaced for the ligand to enter. Docking programs handle water crudely, if at all.
Flexibility. Proteins are not rigid structures. They breathe, flex, and shift conformations. The crystal structure we use for docking is a single snapshot of a dynamic entity. The real binding event involves the protein changing shape to accommodate the ligand — what biochemists call induced fit. Standard docking treats the protein as largely rigid.
Entropy. When a flexible molecule binds to a protein, it loses conformational freedom — it becomes locked in one shape. This loss of entropy has an energetic cost that docking scores don’t fully capture.
Cellular context. A molecule might bind beautifully to a receptor in a docking calculation but never reach that receptor in a living cell. It might get metabolized first. It might not cross the cell membrane. It might get intercepted by a different protein entirely.
These limitations don’t invalidate docking — they define its place in the scientific process. Docking generates hypotheses. It says: “This molecule is predicted to bind this receptor with this affinity. Now go test it.” The lab work — binding assays, cell-based experiments, animal studies, and eventually human trials — is what turns a prediction into a conclusion.
For our coffee diterpenes, the docking predictions are compelling. They align with existing experimental evidence. The scores are strong, the poses are convergent, and the ligand efficiencies are excellent. But they remain predictions until validated by experimental work. I say this not to diminish the results, but to place them honestly within the framework of how science actually works. Computational models generate hypotheses, not conclusions.
Interactive: Caffeine bound to Adenosine A2A Receptor
Rotate, zoom, and explore the 3D structure. Caffeine (green sticks) sits in the orthosteric binding pocket.
Let me distill everything in this chapter down to a single, careful statement.
The molecules in your unfiltered coffee — cafestol and kahweol, those diterpenes that survive in French press, Turkish coffee, and espresso but get caught by paper filters — are predicted to bind the farnesoid X receptor with pharmaceutical-grade affinity. FXR is the nuclear receptor that serves as a master switch for bile acid and cholesterol metabolism. Our computational models predict that these molecules fit into FXR’s binding pocket as snugly as drugs specifically designed for that purpose.
They’re also predicted to interact with other receptors in the same metabolic network — LXR-β, HMGCR, CYP7A1 — with moderate but meaningful affinity. The picture that emerges is of multi-target molecules that touch several points in the cholesterol regulation pathway simultaneously.
That’s not a health claim. It’s a testable prediction from computational science. It’s a hypothesis about why unfiltered coffee raises cholesterol — not just that it does, which epidemiologists established decades ago, but how, at the level of atoms fitting into protein cavities.
The validated caffeine-A2A control gives us confidence that our tools are producing sensible results. The quality metrics — pose convergence, ligand efficiency — tell us the predictions are robust. And the consistency between the docking results and decades of clinical observation tells us we’re probably looking in the right direction.
But probably is not certainly. And a single molecule talking to a single receptor, no matter how tightly, is still a conversation between two. In the next chapter, we zoom out — way out — and watch all fifteen bioactive compounds talking to all their predicted targets at once. Not a conversation. A network. A city of molecular interactions happening simultaneously in the twenty minutes between your first sip and the moment you feel awake.
That’s where the real complexity begins. And honestly? It’s where I started losing sleep for different reasons entirely.
Next: Chapter 5 — “The Network: When Molecules Talk to Each Other”
How grapefruit juice makes your caffeine last longer — real pharmacology in your kitchen
You'll Need
- Two mornings to test
- Your usual coffee
- A glass of grapefruit juice
- Notebook
- Timer
Do This
- Day 1 (control): Drink your normal coffee. Note when you feel peak alertness and when the effect fades. Record the times.
- Day 2 (test): Drink a glass of grapefruit juice 30 minutes BEFORE your coffee. Same coffee, same amount.
- Note peak alertness and when the effect fades.
- Compare your two timelines.
What's Happening
Grapefruit contains compounds called furanocoumarins that partially disable the liver enzyme (CYP1A2) responsible for breaking down caffeine. With that enzyme slowed, caffeine lingers longer in your bloodstream — its half-life stretches out. You are not imagining the stronger buzz. You have just performed a real drug-interaction experiment, the same one pharmacology students study in class.
“Caffeine gets the attention. But the polyphenols are doing something far more complex — a network, not solo agents. What I found when I mapped it surprised even me.”